
The bar graph shows the number of cDNAs that identified each crosslink site. (F) Crosslink sites of TIA1 and TIAL1 in the 3′ UTR of the MYC gene. The Pearson correlation coefficient (r) between the TIA1 and TIAL1 cDNA counts in the clusters is shown. 
The darkness of contours increases with the number of clusters. (E) Contour plot comparing TIA1 and TIAL1 cDNA counts in the 46,970 crosslink clusters. Percentage of crosslink clusters with a given cDNA count in TIA1 iCLIP that were also identified in TIAL1 iCLIP is shown. (D) Reproducibility of TIA1 and TIAL1 crosslink clusters. The sequences of the two most enriched pentamers and the Pearson correlation coefficient (r) between the TIA1 and TIAL1 z scores are shown. (C) Pentamer z scores at the 21 nt sequence surrounding crosslink sites (−10 nt to +10 nt) are shown for TIA1 and TIAL1 iCLIP. (B) The fold enrichment of average cDNA density from TIA1 and TIAL1 iCLIP in different types of RNAs relative to the average cDNA density in the whole genome. (A) The percentage of cDNAs from TIA1 and TIAL1 iCLIP that mapped to different types of RNAs. TIA1 and TIAL1 crosslink to the same positions in human RNAs. The exon and intron positions of the two isoforms of CLIP4 mRNA are shown at the bottom. 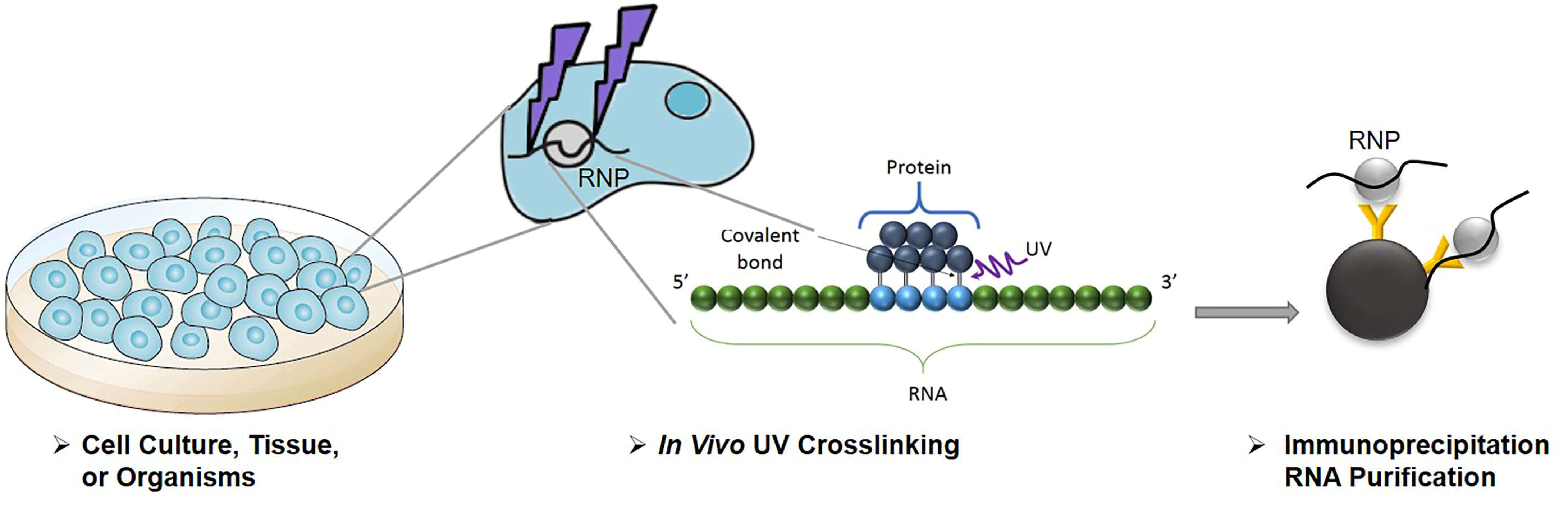
Pre-mRNA sequence is shown below with crosslink nucleotides in red. Below, the bar graphs show the cDNA count (number of cDNAs at each crosslink site). The cDNA positions are colour-coded for three replicate TIA1 and TIAL1 experiments, and the random barcode (shown on the left) is used to identify unique iCLIP cDNAs (number in brackets indicates the number of corresponding PCR duplicates). (C) TIA1 and TIAL1 crosslink to uridine tracts downstream of the alternative 5′ splice sites in the CLIP4 gene. The Western blots below the autoradiograms show the input lysate used for each immunoprecipitation. High and low RNase concentrations were used and protein G beads were used as a control. Immunoprecipitation was performed with either anti-TIA1 or anti-TIAL1 antibody using lysate from UV-crosslinked HeLa cells, cells transfected with TIA1 or TIAL1, TIA1/TIAL1 KD cells, or non-crosslinked cells. ICLIP identifies the TIA1 and TIAL1 crosslink sites with nucleotide resolution.Īutoradiogram of 32P-labelled RNA crosslinked to TIA1 (A) or TIAL1 (B) in HeLa cells. Thus, our findings indicate that changes in splicing kinetics could mediate the distal regulation of alternative splicing. These data are consistent with a model where TIA proteins shorten the time available for definition of an alternative exon by enhancing recognition of the preceding 5' splice site. Using transcriptome-wide high-resolution mapping of TIA-RNA interactions we evaluated the distal splicing effects of TIA proteins. Surprisingly, TIA binding at 5' splice sites silenced distal cassette and variable-length exons without binding in proximity to the regulated alternative 3' splice sites. 
The predictions were validated in an unbiased manner using splice-junction microarrays, RT-PCR, and minigene constructs, which showed that TIA proteins maintain splicing fidelity and regulate alternative splicing by binding exclusively downstream of 5' splice sites. Binding downstream of 5' splice sites was used to predict the effects of TIA proteins in enhancing inclusion of proximal exons and silencing inclusion of distal exons. We used UV-crosslinking and immunoprecipitation (iCLIP) to find that TIA1 and TIAL1 bind at the same positions on human RNAs. 
However, effects of TIA proteins on splicing of distal exons have not yet been explored. T-cell intracellular antigen 1 (TIA1) and TIA1-like 1 (TIAL1) locally enhance exon inclusion by recruiting U1 snRNP to 5' splice sites. The regulation of alternative splicing involves interactions between RNA-binding proteins and pre-mRNA positions close to the splice sites. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
